|
To mix those together in one series fells strange to me. This might be a bug.įrom a user point of view the additional option to open an label images using a distinct way would be more more convenient in my opinion, because the user either wants to see the real image data or the attachment. Will the label image always be the last inside the whole series?Īnd currently this label image (BioFormats 5.2.4) returns an empty image with the wrong XY dimensions. 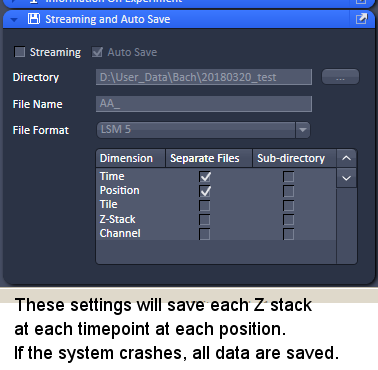
2 Wells with 3x3 Tile Region each -> 18 Series? (by the way, the 2nd approach leads to a error -> separate issue).in an up-to-date Fiji installation, the following exception is thrown: (Fiji Is Just) ImageJ. features of the ZEN core package or simply install it as a viewer for your CZI files. 2 Wells with 9 Positions each -> 18 Series When trying to open a CZI file via File > Import > Image. Olympus viewer software allows you to open.What also puzzle me is the fact that this approach does not really reflect the difference between Therefore any automatic script that used to work will now have to deal with this one image inside the series, that is "different". Now this will open the attachment as well and its dimension will be different from all the other image series. This used to be 96 series, which opened as a stack of 96 images usually, when the Open All option was checked. Imagine one acquires images from a wellplate, e.g. If you are an interested developer you can register for free to get access to the CZI documentation.This sounds fine, but I still think there are some open questions here. Instead, ZEISS supports open software projects and communities with developer-level access, documentation and SDKs for selected microscopy products. Please understand that ZEISS Microscopy cannot provide any kind of user-level assistance for these applications. In addition to this file, the Data Files category includes 1326 related files. What I want to do is to measure their columnarity, i.e.1 answer Top answer: Select Image > Stacks > Orthogonal Views (Mac shortcut is cmd + shift + H) this will present the YZ and XZ views to the right and bottom of the XY view. The file in the CZI format belongs to the Data Files category. 
CZI image and meta-data import is supported through the Bio-Formats plug-in of the Open Microscopy Environment. czi file, which is an image of some cells. CZI files are used for generating video sequences and performing medical research. It contains image stacks, time-lapse series, and tile images captured from a Carl Zeiss microscope. Precompiled installation files of these open source imaging programs are available for Mac OS X and a number of Linux distributions, as well as a multitude of plug-ins for analysis and processing. A CZI file is a microscope image data file created in the Carl Zeiss CZI format. 
If you wish to view and process data that has been created in this format on non-Windows plattforms we recommend to visit the project homepages of ImageJ and the ImageJ-based Fiji software package. ImageJ can directly open TIFF files that have been compressed this way. Images created with ZEN software and ZEISS microscopy systems are saved in the fully documented file format CZI. Some file formats are suitable for data to analyze, others for data only to. Youll want to use the settings: Type 'Positions from File' Order. The Stitching plugins, conversely, are very configurable and use Bio-Formats internally to read the metadata, which is then used to perform the stitching. To support you on other operating systems such as Mac OS X or Linux, ZEISS offers documentation for interoperability and in addition established cooperations with partner projects and companies. The Bio-Formats stitching is an extremely limited feature which does not make full use of the stage position metadata. ZEN software and microscopy imaging workstations by ZEISS are designed to run on 64bit Microsoft Windows platforms.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
